Getting started with #rturf
I started using R in 2011. I’ve read some books, done a lot of Google searches, read a lot of questions and answers that turned up on stackoverflow after executing those searches, and read a lot of blog posts and package vignettes and manuals and documentation. There’s been lots of self-study.
I still consider myself an amateur at this; I’m certainly still a learner. But since I’ve been using this software for so long, along with some other software that works well with R, I suppose I may have learned a few things by now that might help others in the turfgrass world who are interested in what they may be able to do with R.
In that spirit, I’m starting this series of #rturf posts. I may do some videos/screencasts to demonstrate a few things in the future, but it’s easier for me to write and get started with blog posts.
When I was in graduate school, I think the first semester of Biometry 601 used Minitab software. Then subsequent statistics classes all used SAS, and I used SigmaPlot to make charts for presentations and articles. After I graduated I tried to continue using SAS and SigmaPlot but eventually found it too difficult to get licenses. A friend—thanks Josh—suggested R, and that was the first time I heard of it. As I looked into R, I realized I’d be able to do everything I wanted to do if I could learn to use that software. I regret that I did not learn about R, and start using it, even sooner in my career.
What would I use R for? That’s the important question. In my case, I had a few things I wanted to use this type of software for. And I think this is what a lot of graduate students and turfgrass scientists would want to do—communication. Data analysis and then presentation of results.
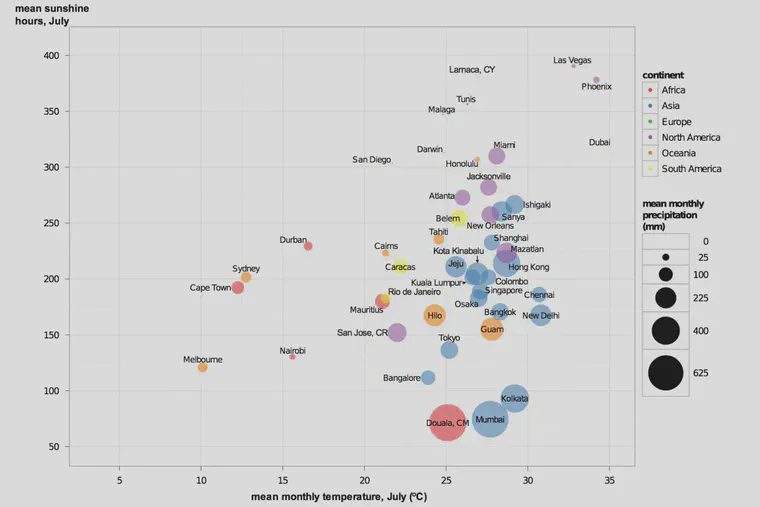
I wanted to make charts, for myself, and for showing in presentations, and for use in articles.
I wanted to produce reports—soil test reports, specifically—that would contain some text and some charts and the report would change when fed a different set of input data.
I wanted to be able to do statisical calculations and data analysis.
I’ve been using R for all those things, and have discovered a lot of other things I can create as well, like Shiny apps and presentation trailers.
In future posts in this series, I’ll share more of the how of R. How to do various things, or how I do various things, that I think might be useful for people working with turfgrass. It may not be a tremendously long series; it may be as I do this that it turns out the readers already know much of what I have to share. We’ll see.
But I want to cover a little more of the why here at the beginning. I was reading the preface to McElreath’s Statistical Rethinking the other day, and he explained why using this type of software is the right way for scientists to do this type of work. I quote here at length from that explanation, because it makes a strong case for why one should make the effort to do things this way.
Programming at the level needed to perform twenty-first century statistical inference is not that complicated, but it is unfamiliar at first. Why not just teach the reader how to do all of this with a point-and-click program? There are big advantages to doing statistics with text commands, rather than pointing and clicking on menus.
Everyone knows that the command line is more powerful. But it also saves you time and fulfills ethical obligations. With a command script, each analysis documents itself, so that years from now you can come back to your analysis and replicate it exactly. You can re-use your old files and send them to colleagues. Pointing and clicking, however, leaves no trail of breadcrumbs. A file with your R commands inside it does. Once you get in the habit of planning, running, and preserving your statistical analyses in this way, it pays for itself many times over. With point-and-click, you pay down the road, rather than only up front. It is also a basic ethical requirement of science that our analyses be fully documented and repeatable. The integrity of peer review and the cumulative progress of research depend upon it. A command line statistical program makes this documentation natural. A point-and-click interface does not. Be ethical.
So we don’t use the command line because we are hardcore or elitist (although we might be). We use the command line because it is better. It is harder at first. Unlike the point-and-click interface, you do have to learn a basic set of commands to get started with a command line interface. However, the ethical and cost saving advantages are worth the inconvenience.
Another example comes to mind, from Andrew Gelman’s blog. He showed a foolish, embarrasing mistake—poll results on a CNN screen display that add to 110% rather than 100%—and mentioned that such a mistake can be prevented when one uses an integrated system. Even better, if one does make a mistake, it can be found and corrected. That’s the ethical bit McElreath was describing.
I could go on and on about the various advantages I’ve found in my own work. But this post is long enough, and I intend to make this rturf category on the blog a series. So I can write more another time, and will start getting into some more practical things with R, in the next installment.